- Home
- Prototypes
- NASFinder
NASFinder
The Network Activity Score Finder (NASFinder) is a web service for topological and functional analysis of sub-networks connecting an omics-determined module.
The web application requires a list of molecules of interest and their experimental data.
Molecules of Interest are those we want to explore their interaction level with the wider surrounding experimental data.
The list of molecules of interest can be determined in many ways including a selection for which we want to verify a hypothesis.
The NASFinder implementation is now available upon request.
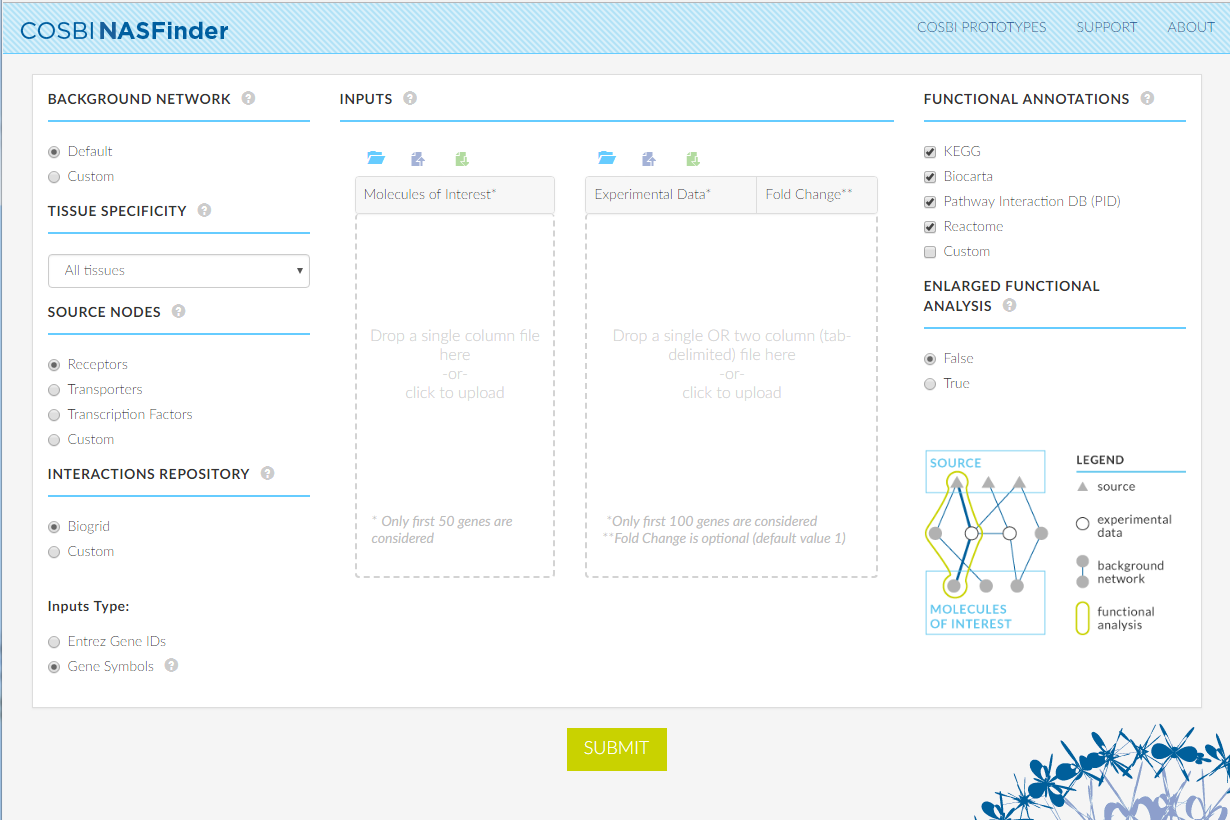
Methods
The identification of active network modules is guided by the desired role for source nodes. Predefined bio-molecular roles available include receptors, transcription factors and transporters.
A spectrum of data, by default from human organism, is integrated in NASFinder, such as:
- the topology of a directed and un-weighted interaction network, called the background network,
- a list of roles for the source nodes to connect to in the background network (e.g. receptors, transporters, or transcription factors),
- repositories of interactions for extending the background network with the user-provided experimental data, and
- functional annotations of reference gene sets for the enrichment phase.
The web server provides the option to customize all of the default data.
Results
The output consists of the network functional analysis result, visualization of omics data sets and network modules.
In addition, web interface provides download options for the network topology and enrichment analysis results.
The network topology is visualized using Cytoscape.js
References
Nassiri, I., Lombardo, R., Lauria, M. et al. Systems view of adipogenesis via novel omics-driven and tissue-specific activity scoring of network functional modules. Sci Rep 6, 28851 (2016). link