Computational analysis of cytokine release following bispecific T-cell engager therapy
- Expertise
- Modeling & simulation
Challenge
Develop a logical model describing the interplay between Bispecific T-cell engager antibodies (BiAbs) the immune system, and tumor cells. Perform in silico simulations to elucidate the factors affecting the cytokine release syndrome (CRS). Identify effectiveness of co-medications to administer with BiAbs molecules to mitigate CRS without affecting the efficacy.
What we did
The logical model was built focusing on the major variables influencing the CRS activation and the tumor cell elimination. The rules governing the interactions were defined based on known biology and published experimental evidence. The syndrome severity was described by the phenotypic readout Combined Cytokine Index CCI used as a local proxy for the CRS observed in the clinical setting. The efficacy of co-meds was assessed by performing a sensitivity analysis of CCI and tumor with respect to model mutations (nodes knock-down or over-activation).
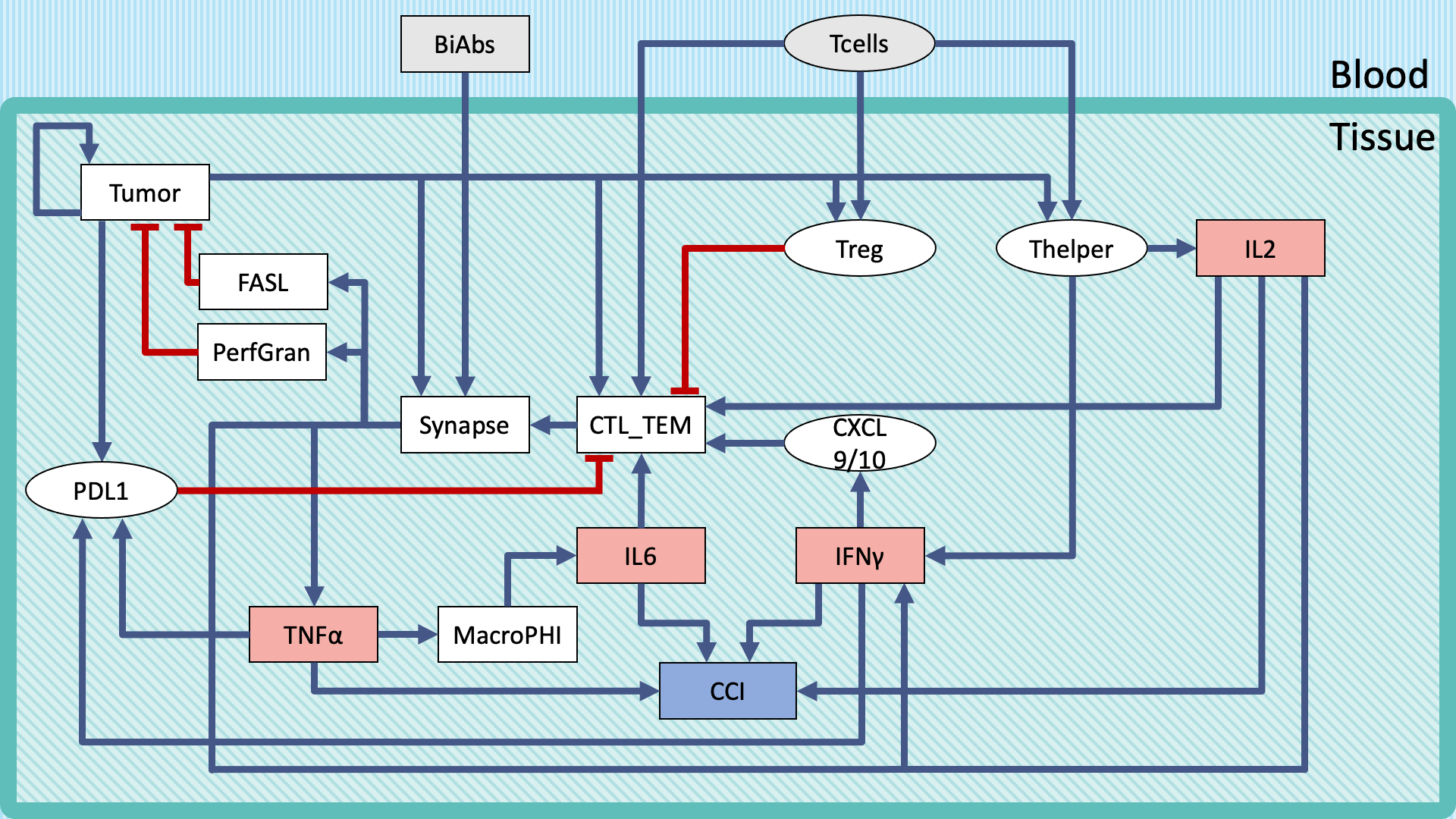
Outcome
The model developed enables a systematic analysis in which is possible to test combinatorially single or multiple co-meds treatments to provide together with BiAbs. The minimal requirement of information to set up the model and the qualitative outcome of the results makes it ideal for exploratory analysis.
Partner

Amgen is committed to unlocking the potential of biology for patients suffering from serious illnesses by discovering, developing, manufacturing and delivering innovative human therapeutics.
https://www.amgen.com/Publications
Selvaggio G, Parolo S, Bora P, Leonardelli L, Harrold J, Mehta K, Rock DA, Marchetti L, Computational Analysis of Cytokine Release Following Bispecific T-Cell Engager Therapy: Applications of a Logic-Based Model. Frontiers in Oncology 12, 2022.
See 10.3389/fonc.2022.818641
Selvaggio G, Parameter free approaches in QSP: modelling the cytokine release following bispecific T-cell engager therapy, Annual meeting of the Society for Mathematical Biology, June 13-17, 2021. http://schedule.smb2021.org/MS06/MFBM-MS06.html